Learning objectives
- Explain how plants and other photoautotrophs create biomass mostly from carbon dioxide in the air.
- Describe the interdependence of the light reactions and carbon fixation reactions
- Explain why occasional cyclic electron flow in the light reactions is required for operation of the Calvin cycle
- Predict how disruptions in the Calvin cycle affect concentrations of key compounds
- Describe the activities and functions of rubisco
- Summarize the inputs and outputs of PSI, PSII, and the Calvin cycle using NADPH, glucose, H2O, O2, CO2, H+ gradients, and ATP
Carbon fixation is neither “dark” nor “light-independent”
You may have previously learned about carbon fixation using the terms dark reactions or light-independent reactions. In Biological Principles,
- we will avoid the term “dark” reactions because these reactions can occur in the light.
- we will also avoid the term “light-independent” reactions because these reactions do depend on the products of the light reactions: ATP and NADPH.
Instead, we will simply call these reactions the process of “carbon fixation” or the Calvin Cycle.
Before we begin, here is a reminder of where we left off: how the light reactions work, and the products they generate which are essential for the Calvin cycle. (The video below has an error starting at 10:15. The high energy electron carrier in chloroplasts is NADPH, not NADH.)
And here is the introduction to where we’re going:
Where do plants get their biomass?
Jan Baptiste van Helmont performed a famous experiment where he planted a willow sapling in a pot with exactly 200 pounds of soil, largely covered except for holes for watering. After 5 years of watering, he removed and weighed the tree and the soil left in the pot, after he had carefully recovered the soil clinging to the roots. He found that the tree’s weight had increased by 75.4 kg, but the soil had lost only 57 g. He concluded, reasonably but erroneously, that most of the mass of the tree came from water.
But water does not contain carbon atoms. The biomass of the tree, consisting largely of carbohydrates (cellulose, lignin, etc.), comes from carbon dioxide:
CO2 + 12 H2O –> C6H12O6 + 6 O2 + 6 H2O
Only the hydrogen atoms from water molecules contribute to the biomass.
This video explains how this process (carbon fixation) works in the Calvin cycle:
How do we know how carbon fixation works?
Here’s how Calvin, Benson, and Bassham worked out carbon fixation. Calvin and colleagues used radioactive 14CO2 (carbon dioxide labeled with 14-carbon) to determine what are the first products of carbon fixation by a culture of algae. Within 10 minutes after injecting 14CO2, many kinds of organic molecules were radioactively labeled. At shorter times, down to a few seconds, they found that the first product of carbon fixation was a 3-carbon sugar, 3-phosphoglycerate (3-PG), labeled at the carboxyl group.
Using ATP and NADPH from the light reactions, 3-PG is reduced to glyceraldehyde-3-phosphate (G3P). Two molecules of G3P can spontaneously form a molecule of glucose (the first, energetically-uphill part of glycolysis running backwards).
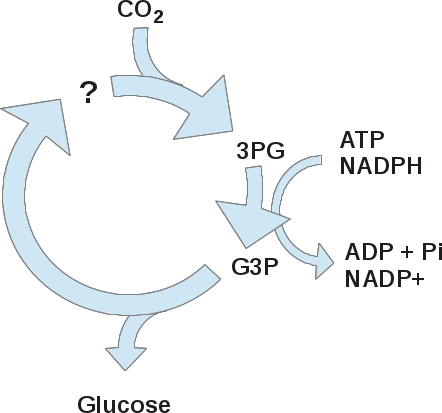
However, they were unable at first to identify a 2-carbon molecule that could bond with CO2 to form 3-PG. They did deduce that this was a cyclic process, because at longer times, the other carbons of 3-PG became labeled. This figure shows a model of their initial findings:
Based on this model, Calvin and colleagues predicted that, in the dark, 3PG would accumulate because no ATP or NADPH would be made, and all other intermediates of the cycle would decrease because they are eventually converted to 3PG. On the other hand, in the absence of CO2, whichever molecule that accepts CO2 would increase, because all the other intermediates in the cycle would be converted to this acceptor compound.
The acceptor compound turned out to be a 5-carbon sugar phosphate, ribulose-1,5-bisphosphate (RuBP). The enzyme rubisco (RuBP carboxylase/oxygenase) attaches CO2 to RuBP to form a highly unstable 6-carbon intermediate that immediately breaks down into 2 3-carbon sugars, 3PG.
You can step through an interactive animation here: https://www.science.smith.edu/departments/Biology/Bio231/calvin.html
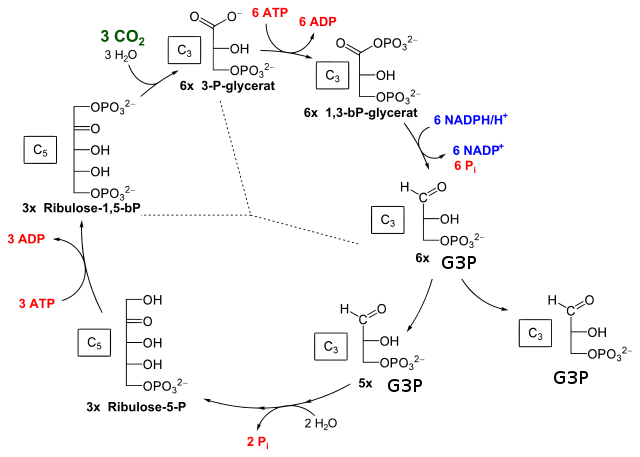
- Every 3 molecules of CO2 fixed by rubisco produces 6 molecules of 3PG.
- Conversion of 6 molecules of 3PG to 6 molecules of G3P consumes 6 ATP and 6 NADPH.
- One molecule of G3P exits the cycle, leaving 5 molecules of G3P (15 carbon atoms)
- The 5 molecules of G3P undergo a series of rearrangements to form 3 molecules of ribulose-5-phosphate (15 carbon atoms)
- The phosphorylation of 3 molecules of ribulose-5-phosphate to restore 3 molecules of ribulose-1,5-bisphosphate consumes 3 more ATPs
- net: 3 CO2 fixed to make 1 molecule of G3P, consuming 6 NADPH and 9 ATP
- 2 molecules of G3P combine to make 1 molecule of glucose
The products of the Calvin cycle are exported from the chloroplast to provide energy for cellular respiration, and organic carbon for biosynthetic pathways to construct all the building blocks for cellular molecules.
If you’d like more information, below is Dr. Choi’s 35-min screencast of the Calvin cycle and C4 photosynthesis (this video includes shorter segments excerpted above)
And the powerpoint slides for the screencast: photosynthesis_carbonfixation
Sustainable Development Goals


UN Sustainable Development Goal (SDG) 14 Life Below Water and SDG 15: Life on Land – The ability of plants and other photoautotrophs to create biomass from carbon dioxide in the air is essential for sustaining life on Earth. Through the process of photosynthesis, these organisms capture light energy from the sun and convert it into chemical energy in the form of carbohydrates, which can be used for growth and reproduction of these organisms and those that eat them. Photosynthesis also releases oxygen into the atmosphere, which is crucial for supporting aerobic respiration in many organisms, including us!



Is the Calvin Cycle part of photosynthesis?
Yes. Photosynthesis is the use of light energy to “fix” (reduce) inorganic carbon to organic carbon, so the Calvin cycle is the pathway that actually does the reduction of carbon dioxide to form carbohydrates.