Learning Objectives
- Define the chromosome theory of inheritance as “genes are located on chromosomes”
- Use phenotypic ratios to determine if genes are sex-linked
- Predict possible offspring types and phenotypic ratios in the case of sex linkage
- Apply pedigree analysis to distinguish between dominant, recessive, and sex-linked traits
- Define linkage as departure from independent assortment
Genes are located on chromosomes
We take it for granted today that DNA is the genetic material, and therefore our genes must be located on chromosomes. But like all facts in science, this idea had to be repeatedly tested and found to be true before it could be accepted as fact. The chromosome theory of inheritance, or the idea that genes are located on chromosomes, was proposed based on experiments by Thomas Hunt Morgan using Drosophila melanogaster, or fruit flies. Drosophila are like humans in that an individual with two X chromosomes is female and an individual with one X and one Y chromosome is male (many organisms have other ways of determining gender).
In Drosophila, normal flies have red eyes. Red eye color is dominant. Morgan discovered a recessive mutation (allele) that caused white eyes. When Morgan mated a red eyed female to a white eyed male, all the progeny had red eyes. This result makes perfect sense with a dominant/recessive inheritance pattern, and here is the Punnett square demonstrating that (x^w = recessive white eye mutant allele; x^W = dominant red eye wild-type allele):
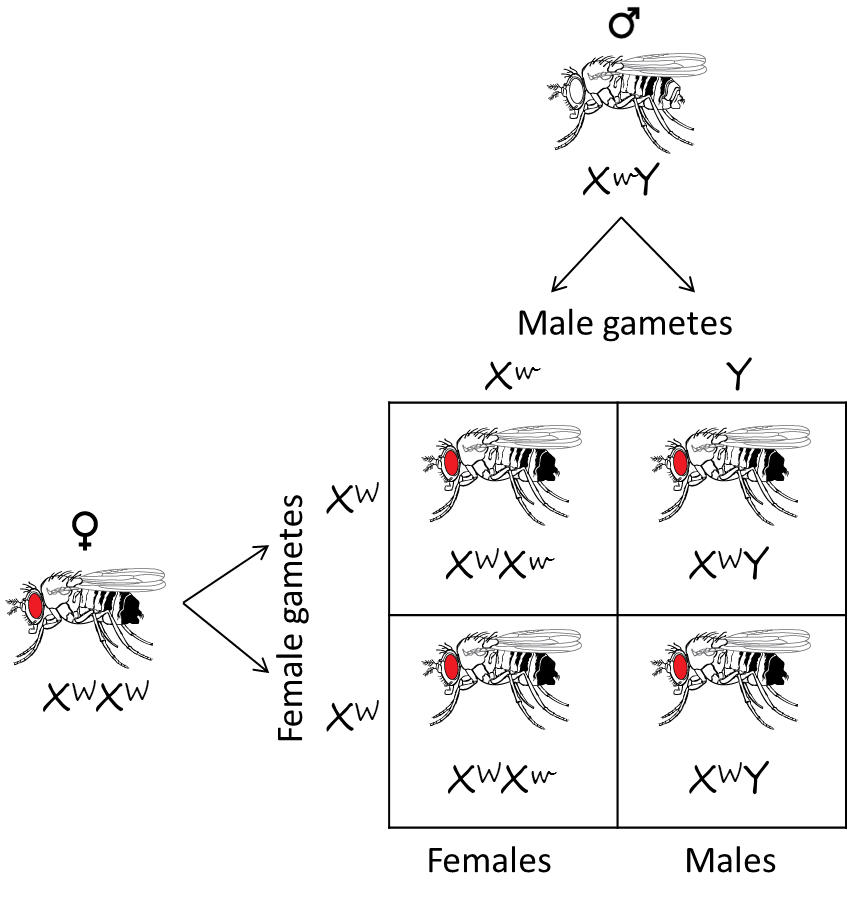
Adapted from OpenStax Biology (http://cnx.org/resources/9ce8757f364f530db58306d982c0dbc52932e235/Figure_12_02_09.jpg)
But Morgan got a surprising result when he made the reciprocal cross, mating white eyed females to red eyed males. Instead of all red eyed progeny, he saw that all the females had red eyes and all the males had white eyes. This result seemed to violate Mendel’s principle of independent assortment, because two different traits (gender and eye color) seemed to be linked. The only way to explain these results was if the gene that caused eye color was located on (linked to) the X chromosome. Here is the Punnett square demonstrating this cross:
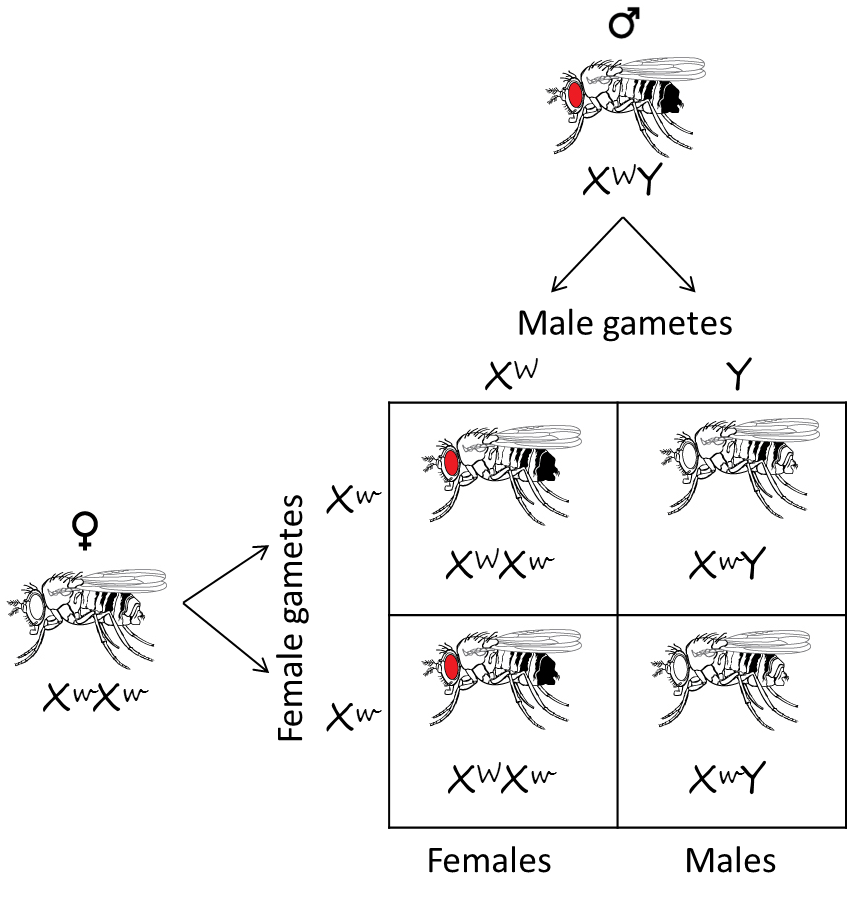
Adapted from OpenStax Biology (http://cnx.org/resources/9ce8757f364f530db58306d982c0dbc52932e235/Figure_12_02_09.jpg)
These results support the chromosome theory of inheritance because the only way to explain them is if the eye color gene is on the X chromosome. This is sex-linkage, or inheritance of genes that are on the sex chromosomes (X and Y). Sex-linked traits show interesting inheritance patterns in part because females have two copies of each X chromosome, but males only have one. This inheritance pattern means that a male with the recessive allele will always show the recessive trait, because he only has one copy of the allele. In contrast, most genes are located on the autosomes, or non sex chromosomes, where both males and females have two copies of each gene. Recall that all the patterns of inheritance observed by Mendel, including the principle of segregation and the principle of independent assortment are explained by the behavior of chromosomes during meiosis. These principles are part of the chromosome theory of inheritance.
Here is a video explaining these experiments and a little on the implications for humans:
In class, we’ll use phenotypic ratios to determine whether genes are sex-linked and predict offspring phenotypes when genes are sex-linked. We’ll also apply this information to analyze human pedigrees.
Linkage
Linkage is inheritance of traits in a pattern that violates Mendel’s principle of independent assortment, the idea that alleles for different traits are segregated into gametes independently. Sex-linkage is a special type of linkage, where traits are linked to sex chromosomes. Genetic linkage occurs when the genes controlling two different traits are located near each other on the same chromosome. The basic idea is that if two genes are on the same chromosome, and you inherit the whole chromosome, then you have to inherit those two genes (and whatever alleles they have) together.
However, this is biology so there is a caveat: the phenomenon of crossing over helps to shuffle the alleles for genes located on the same chromosome. A crossover event between the locations of two genes on a chromosome results in genetic recombination, or new combinations of alleles on a chromosome.

Crossing over between genes A and B results in recombinant chromosomes with new allele combinations a, b and A, B, in addition to the original parental combinations A, b and a, B. Image from Wikimedia by user Abbyprovenzano, with CC-BY-SA-3.0 license.
Crossing over occurs during meiotic prophase I, when the homologous chromosomes align and synapse, and results in physically swapping genetic material (DNA) between non-sister chromatids of the paired homologous chromosomes. Because crossing over occurs randomly along the chromosome, the closer two genes are physically located to each other on a chromosome, the less likely that a crossover will occur between them. Conversely, the farther apart two genes are located from each other along the chromosome, the more likely they are to be swapped with the alleles on the homologous chromosome. The image below illustrates this idea:
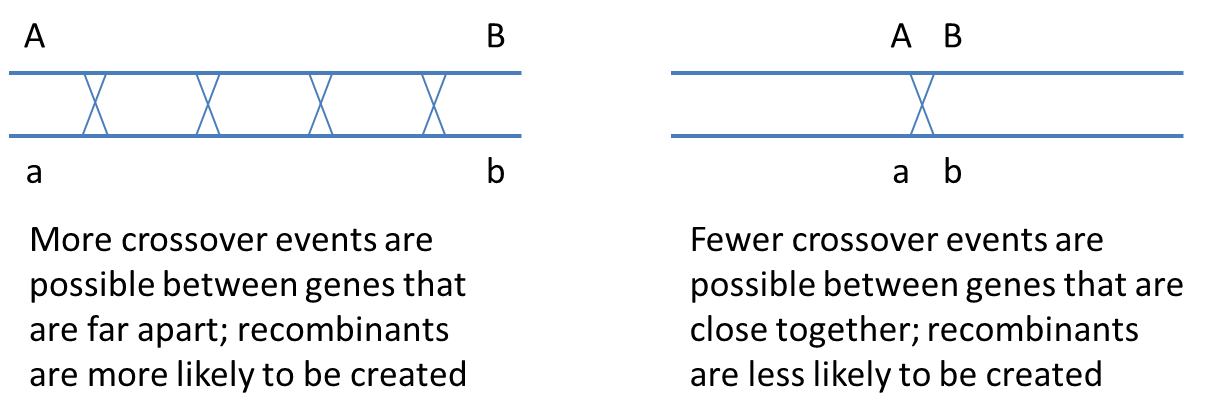
It may be surprising to realize that two genes on the same chromosome will assort independently (like genes located on separate chromosomes) if they are far enough apart that a crossover almost always occurs between them, producing 50% recombinants (because crossing over involves only two of the 4 chromatids in a synapsed pair of homologous chromosomes, the maximum recombination frequency is 50%).
The video below walks through linkage as a violation of independent assortment and explains how crossing over breaks linkage. Note this video uses an incomplete definition of linkage: linkage occurs when two genes are located close together on the same chromosome and thus tend to be inherited together. It is not sufficient for genes to be on the same chromosome to be linked; they also have to be close enough together that crossing over between them is a relatively rare event.
Simple rules for pedigree analysis
We can’t ask different people to mate and produce lots of offspring so we can test inheritance patterns in humans. Instead we rely on pedigree analysis to infer inheritance patterns. Here is a sample pedigree which explains how to read pedigrees:
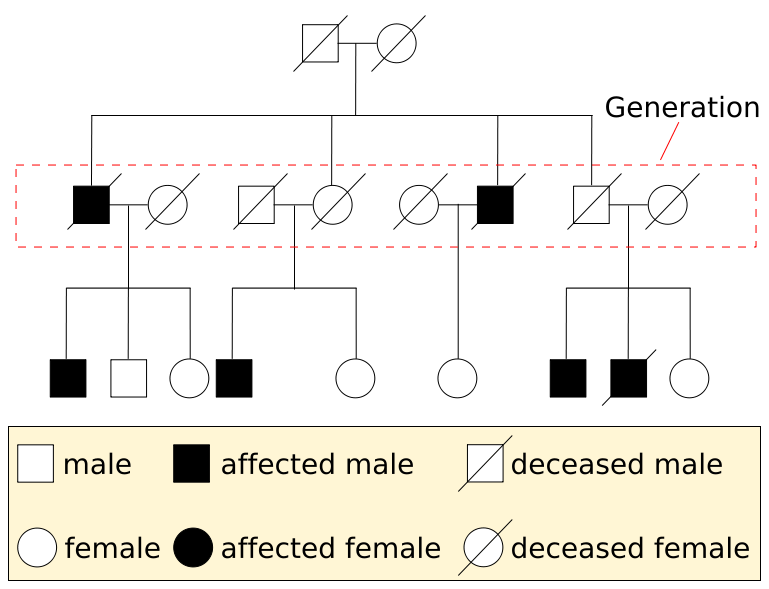
Source: Wikimedia Commons (https://upload.wikimedia.org/wikipedia/commons/thumb/9/9c/Pedigree-chart-example.svg/769px-Pedigree-chart-example.svg.png)
The simple rules for pedigree analysis are:
- Autosomal recessive
- affects males and females equally
- both parents must carry allele
- parents may not display trait (carriers)
- ~1/4 of children affected (if both parents are carriers)
- Autosomal dominant
- affects males and females equally
- only one parent must carry allele
- if child displays trait, at least one parent must also display trait
- ~1/2 of children affected (if one parent displays trait)
- X-linked recessive
- typically affects only males
- affected male passes allele to daughters, not to sons
- trait skips a generation
In class, we will practice using these rules to determine the inheritance patterns of traits in different pedigrees.

https://xkcd.com/634/
Powerpoint slides with animated illustration of chromosome movements in mitosis and meiosis, to accompany the Nash case: MollyNashMitosisMeiosisAnimations

UN Sustainable Development Goal (SDG) 3: Good Health and Well-being – Understanding the chromosome theory of inheritance is important in identifying the genetic basis of diseases and developing treatments for genetic disorders. Because genes are not inherited in the same patterns (i.e. pedigrees differ whether a trait is dominant, recessive, or sex-linked), applying pedigree analysis can help us better understand new diseases and provide targeted support in specific communities for equitable access to healthcare.


I removed the video on crossing over and recombination because it was way too technical, but if you want to delve into it, here it is: https://www.youtube.com/watch?v=BhJf9MHHmc4
The case of Lisa and Jack Nash, who conceived a child by in vitro fertilization and pre-implantation genetic diagnosis to serve as a tissue donor for their daughter who had Fanconi anemia, illustrates both linkage of HLA genes and independent assortment of the HLA genes with respect to the Fanconi anemia locus. The original research paper is here: http://jama.jamanetwork.com/article.aspx?articleid=193944