Learning objectives:
- Know the functions of the three types of RNA
- Describe the process and factors of transcription and predict outcomes if one factor is missing
- Predict the RNA transcribed from a DNA sequence identified as either the template strand or the coding strand
- Use the genetic code to predict the protein amino acid sequence translated from an mRNA sequence
- Describe the process and factors of translation and predict outcomes if one factor is missing
- Describe the DNA sequence motifs and proteins required to initiate transcription
- Predict the likely effects of mutations in DNA on protein amino acid sequence, structure and function
The Central Dogma
Francis Crick coined this term to describe the flow of information from nucleic acid to protein. Information encoded in DNA is transcribed to RNA, and RNA is translated to a linear sequence of amino acids in protein. Although information can flow reversibly between DNA and RNA via transcription and reverse transcription, no mechanism has yet been found for alterations in protein amino acid sequence to somehow cause a corresponding change in the RNA or DNA.
Transcription: DNA to RNA
- The enzyme RNA polymerase reads the template strand of DNA and synthesizes an RNA molecule whose bases are complementary to the template strand of DNA.
- RNA is synthesized 5′ –> 3′; RNA polymerase reads the template strand of DNA 3′ –> 5′.
- Because the mRNA is complementary to the template strand, the sequence of bases in mRNA is the same as the sequence of bases in the “coding” strand of DNA, except that RNA has uracil (U) instead of thymine (T).
- RNA polymerases in both prokaryotes and eukaryotes depend on DNA-binding proteins, called transcription factors, to bind to special sequence motifs in the DNA called promoters. Transcription cannot start until an RNA polymerase is bound to the promoter.
- Transcription factors recruit RNA polymerase to bind to the promoter sequence and begin transcription just “downstream” of the promoter.
Prokaryotes: transcription and translation are coupled
In prokaryotes, ribosomes begin to translate even while the mRNA is still being transcribed. This can occur because there is no nuclear membrane separating the DNA from the cytoplasm.
Eukaryotes: transcription and translation are separated in space and time, and nuclear pre-mRNA undergoes processing to become mature mRNA
In eukaryotes transcription occurs in the nucleus, whereas translation occurs outside the nucleus, in the cytoplasm by free cytoplasmic ribosomes or ribosomes docked to the ER.
The RNA transcribed from a protein-coding gene in the nucleus is called the “pre-mRNA” or primary mRNA transcript. Pre-mRNA has to undergo at least two, and usually 3, processing steps before they can be exported to the cytoplasm as mature mRNA. These are, in order:
- The 5′ end of the pre-mRNA is modified by the covalent attachment of a 7-methylG nucleotide, called the 5′-cap. The 5′ cap is required for eukaryotic ribosomes to initiate translation.
- For genes with introns (most genes in multicellular eukaryotic organisms), any non-protein coding sequences called introns are removed by RNA splicing, leaving just the exons.
- The 3- end of the pre-mRNA is modified by the addition of hundreds of adenine nucleotides, called the polyA tail. The polyA tail is important for nuclear export, mRNA stability, and translation.
All of these processing steps actually happen while the mRNA is being transcribed; that is, they occur co-transcriptionally. So in reality, a full-length “pre-mRNA” never actually exists.

- Eukaryotic pre-mRNAs are processed in the nucleus by adding a 5′ cap, 3′ polyA tail, and removal of introns via RNA splicing to create a mature mRNA consisting only of exons, ready for export to the cytoplasm for translation. From https://www.biology.arizona.edu/molecular_bio/problem_sets/mol_genetics_of_eukaryotes/03t.html
Molecular animation of transcription and mRNA processing:
BioFlix Transcription
Translation: RNA to Protein
Translating a sequence of bases in the RNA to a sequence of amino acids in proteins requires 3 major components: the messenger RNA (mRNA), ribosomes, and transfer RNAs (tRNAs).
mRNAs are transcribed from protein-coding genes.
Ribosomes are large assemblies of ribosomal RNA molecules (rRNAs) and dozens of proteins. When they are not in the process of translating mRNA into protein, they fall apart into the small subunit and large subunit, each consisting of a rRNA and numerous proteins. When the structures of prokaryotic ribosomes were determined at high resolution, researchers were astonished to discover that the catalytic site for the peptidyl-transfer reaction (attaching new amino acids to the growing polypeptide chain) consists entirely of rRNA. Thus the ribosome is actually an immense ribozyme, a catalytic RNA molecule stabilized by numerous proteins. The protein components provide structural support but are not involved in the protein synthesis reactions.
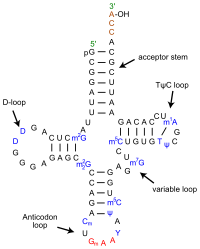
tRNAs match the amino acid to the codon in the mRNA. The bases in the anticodon loop are complementary to the bases in an mRNA codon. The 3′ end of the tRNA is bound to the appropriate amino acid that matches the anticodon. Cells have a family of enzymes, called amino-acyl tRNA synthetases, that recognize the various tRNAs and “charge” them by attaching the correct amino acid.
Translation begins near the 5′ end of the mRNA, with the ribosomal small subunit and a special initiator tRNA carrying the amino acid methionine. In most cases, translation begins at the AUG triplet closest to the 5′ end of the mRNA. The large ribosomal subunit then docks and translation begins. The ribosome moves along the mRNA 3 bases at a time, and new tRNAs whose antibodons are complementary to the mRNA codons arrive with their matching amino acids. A peptide bond forms, the ribosome moves another 3 bases, the empty tRNA is ejected to make room for a new amino-acyl tRNA.
The polypeptide chain that is made also has directionality; one end has a free amino group and the other end of the chain has a free carboxyl group. These are called the N-terminus and the C-terminus, respectively. New amino acids are added only to a free carboxyl end, so polypeptide chains grow from the N-terminus to the C-terminus.
Watch the molecular animation of translation here:
BioFlix Translation
The Genetic Code
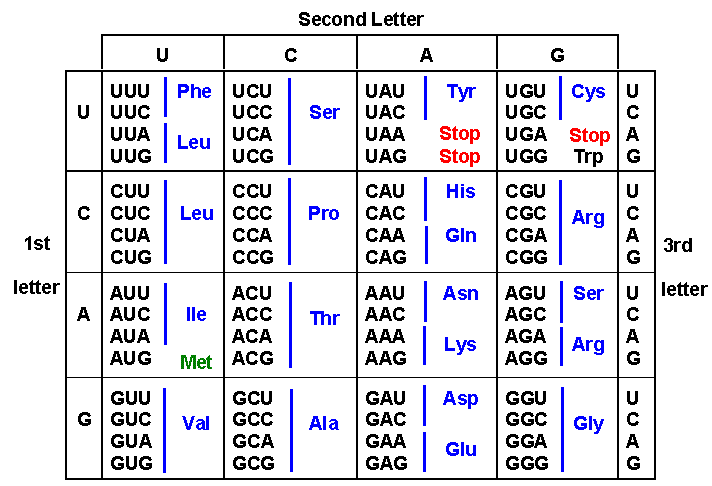
The genetic code is used by all living organisms, whether Archaea, Bacteria or Eukarya, with only minor modifications in the mitochondria of a relatively few species. The code is “degenerate”, because many amino acids are specified by 2, 3, 4 or 6 different codons. See above that phenylalanine (Phe) is coded for by UUU and UUC, while leucine (Leu) is coded for by UUA, UUG, CUU, CUC, CUA, and CUG.
Mutations can have vastly different effects depending on where they occur
If we consider just single nucleotide changes (substitutions, deletions or insertions of single bases), these can have very different consequences depending on whether they occur in the gene.
DNA base substitutions often have no effect if they change the 3rd base in the codon. For example, changing GAG to GAA has no effect on the protein because both codons specify alanine. Such “silent” mutations are called “synonymous” mutations.
Other base substitutions in the first or 2nd position will cause amino acid changes; these are “nonsynonymous” mutations.
Even among nonsynonymous mutations, the exact amino acid change matters. A change of one hydrophobic amino acid to another hydrophobic amino acid will be less disruptive to the structure of the protein than a change of a hydrophobic amino acid to a polar or charged amino acid. Finally, some parts of a protein are more important than others, such as the catalytic site of enzymes, or sites that bind other proteins, DNA, or regulatory molecules.
Insertions or deletions (“indels”) of single nucleotides cause a change in the reading of all downstream codons; they are shifted by one base. Such “frameshift” mutations will alter most or all amino acids downstream (towards the 3′ end of the mRNA, towards the C-terminus of the protein) of the mutation.
Dr. Jung Choi’s video lecture on this topic, in one 32-min chunk (until I find time to split it)
And his slide set:
B1510_module4-6_DNA_to_protein_2011